Protein Docking...the old fashion way
In this post, I revisit a project I worked on as an undergraduate for AMS 548: Optimization Techniques in Biomolecular Simulations at Stony Brook University, taught by Dima Kozakov, back in Spring of 2021. At the time, I was an ambitious undergraduate trying to fulfill requirements for a Chemistry minor alongside my Biochemistry degree, which meant taking a few courses outside my formal curriculum. While browsing options, this course immediately stood out.
Dima is an excellent instructor and a leading researcher in protein-protein docking. His lab develops widely used docking software, so learning directly from someone with deep expertise in the field was both instructive and motivating. Upon doing this write-up, I learn he moved to UT Austin. Below is the official course description:
This practical hands-on course teaches basic techniques for building mathematical models, algorithms, and software for biomolecular simulations of macromolecular interactions. Topics include the basics of statistical mechanics and its connection to sampling algorithms; the origin of and approximations for molecular force computations; the geometry of molecular configuration search spaces and multidimensional optimization; and fundamentals of software development and high-performance computing (HPC). During the course, students develop a multiscale approach to modeling protein-protein interactions from the ground up.
Preface
The course focused on state-of-the-art techniques as they existed in 2021. Fast Fourier Transform (FFT)-based methods for protein docking were already well established, but Kozakov’s innovation was the use of eigenvalue-eigenvector decompositions of pairwise interaction potentials. This reformulation allowed interaction terms to be evaluated efficiently using FFT-based correlations. In the PIPER framework, this effectively introduces a coarse-grained representation of the interaction energy landscape.
Aside: this was a very different era, well before AlphaFold-based docking “hacks” and modern diffusion models changed the field. It raises the question: do fundamental physics-based approaches still matter for protein docking?…
The Project
For the course capstone, we worked with a deliberately simple model. The goal was to develop a program that performs Monte Carlo protein docking using rigid-body representations. While highly simplified, this setup forced us to confront the computational and physical challenges of docking, sampling, scoring, and optimization, without relying on black-box tools (or even LLMs during that time period).
In hindsight, revisiting this project has been a useful reminder of how much insight you can gain from building things the “old-fashioned” way.
Relevant Papers
- PIPER: An FFT-based protein docking program with pairwise potentials
- Modeling Protein-Protein and Protein-Ligand Interactions by the ClusPro Team in CASP16
Why this approach?
When I signed up for this course in November 2020, there wasn’t much buzz yet around AlphaFold, and the role of machine learning in biophysics still felt far away. With that context, it’s easy to overlook traditional docking, but physics-based scoring is still appealing because it’s built from explicit interaction terms rather than learned correlations. That said, these methods are limited by their assumptions (rigid bodies, simplified potentials) and by sampling: if the search can’t efficiently explore relevant configurations, the score won’t help. Recent work from the Kozakov/Vajda labs reflects this reality by combining generative models with more traditional approaches like molecular dynamics (MD) and the PIPER/ClusPro framework. A major concern for purely data driven models is out of distribution behavior when the models encounter interaction patterns that weren’t well represented in training.
Notes & observations
Scoring function
In this project, I implemented a simple Coulomb-like electrostatics term by looping over every receptor-ligand atom pair, reading each atom’s charge from the parameter set, and accumulating a score proportional to:
\[E_{\text{coul}} \propto \sum_{i \in R} \sum_{j \in L} \frac{q_i q_j}{d_{ij}^2}\]I also computed energy gradients (with respect to atomic coordinates) to estimate the direction of the electrostatic force.
For docking acceptance/rejection, I used Dima’s complex_energy routine (energy.h). It evaluates pairwise interactions and wraps them with basic clash/contact handling. That scalar score is what the Monte Carlo loop uses to accept or reject a proposed move.
The pairwise potential term used there is:
\[E_{\text{pair}} = \sum_{i \in A^{*}} \sum_{j \in B^{*}} \mathbf{1}\!\left(r_1^2 \le d_{ij}^2 < r_2^2\right)\, e_{ij}\] \[e_{ij} = \sum_{m=0}^{k-1} \lambda_m\, X_{m,s_i}\, X_{m,s_j}\] \[d_{ij}^2 = (x_i-x_j)^2 + (y_i-y_j)^2 + (z_i-z_j)^2, \qquad s_i = \operatorname{subatoms}(\mathrm{atom\_typen}_i)\]Where:
- $(r_1, r_2, k, \lambda_m, X)$ come from
prms(prms.h). - $A^{}$ and $B^{}$ include all atoms unless
only_sab=1, in which case only atoms withsa != 0are used. - This is the function declared as
pairwise_potential_energy(...)inenergy.h.
In complex_energy (the one used in docking), this pairwise term is combined as:
Note: PIPER accelerates these pairwise potentials via an eigenvalue decomposition and FFT-based correlations, but our course project did not implement that acceleration; we evaluated the pairwise term directly.
The assignment had a couple of parts (I do not have the exact prompt, but this is what I remember):
- Q1a. Compute the center-of-mass (CoM) shift vector between receptor and ligand, translate one structure by half that shift, and write the translated structure to file.
- Q1b. Compute the total Coulombic interaction energy between receptor and ligand atoms using charges from the parameter file.
- Q1c. Compute the analytic gradient of the Coulombic energy with respect to Cartesian coordinates $(F_x, F_y, F_z)$.
- Q1d. Compute the same gradient numerically using finite differences $(\Delta = 0.01)$ and compare against Q1c.
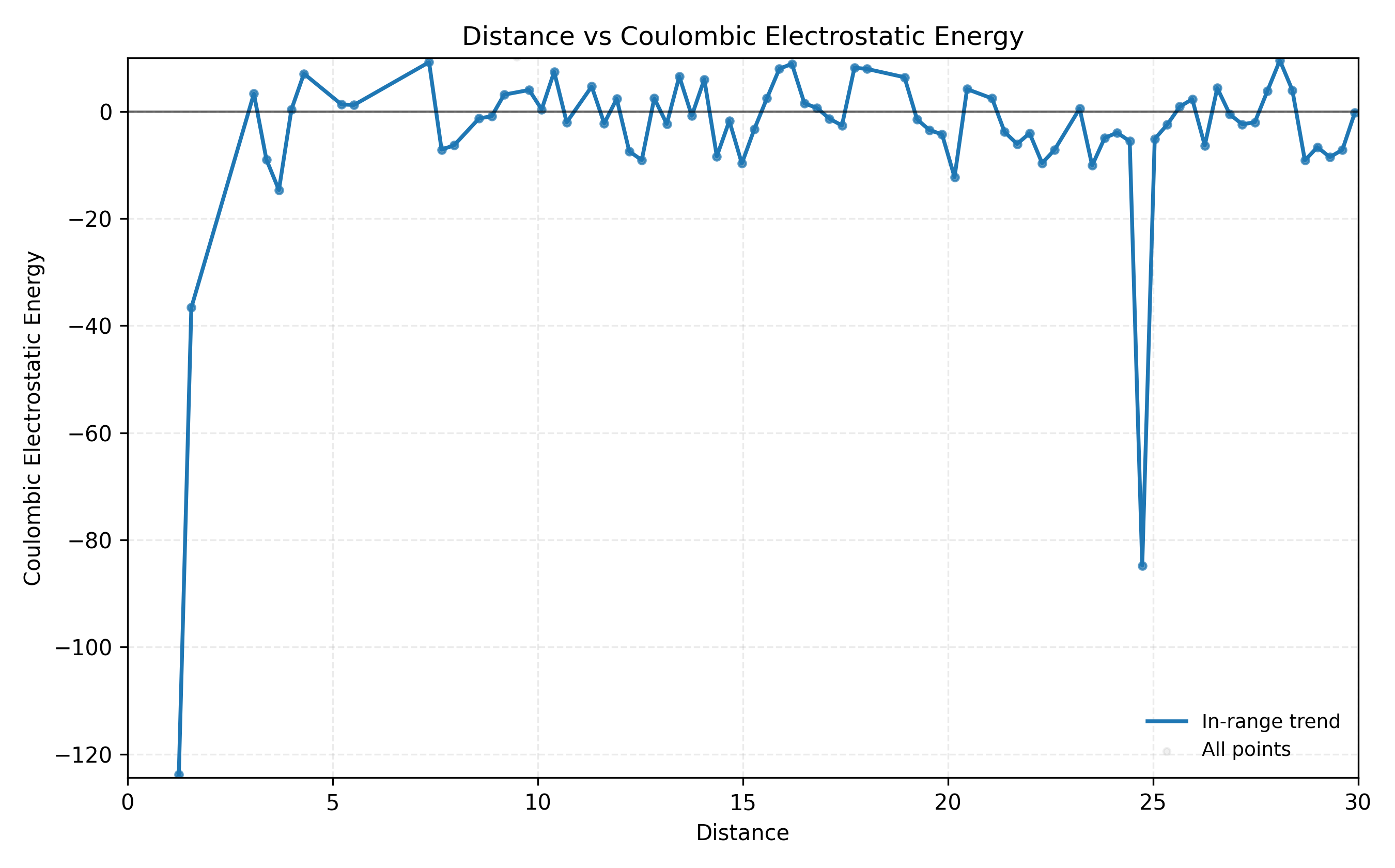
- Q1e. Iteratively translate a structure and report how Coulombic energy changes as CoM distance changes.
- Q2a. Estimate $\pi$ with a Monte Carlo simulation.
- Q2b. Use a 1D Metropolis Monte Carlo method to minimize the toy energy function:
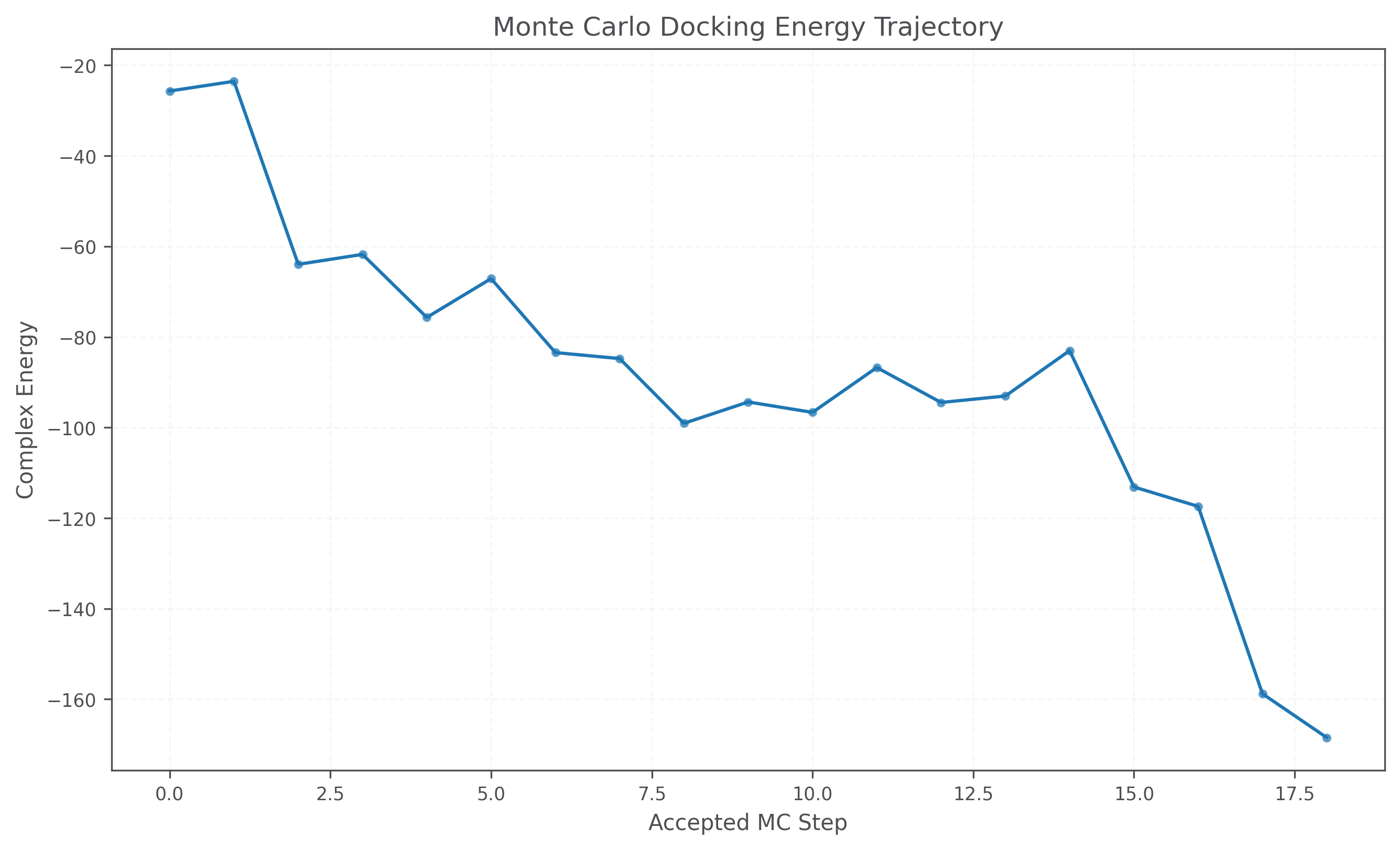
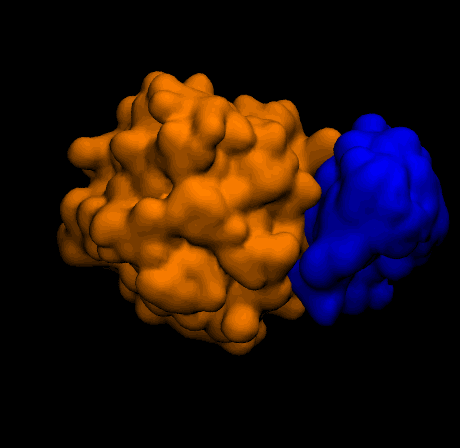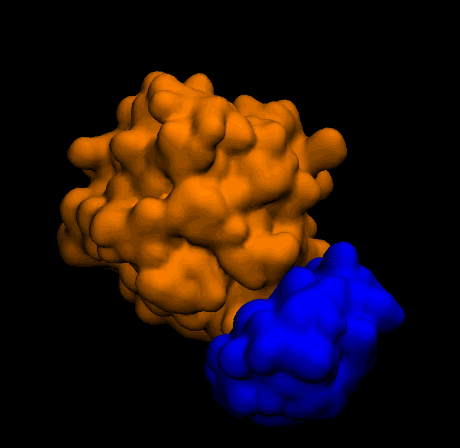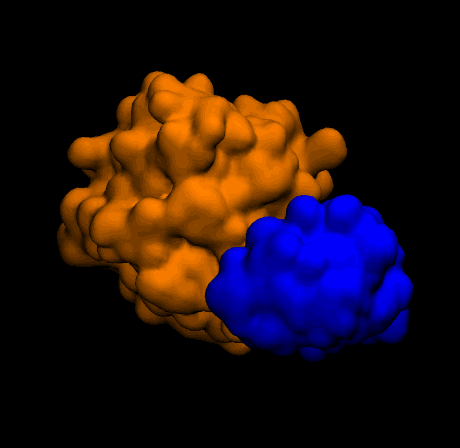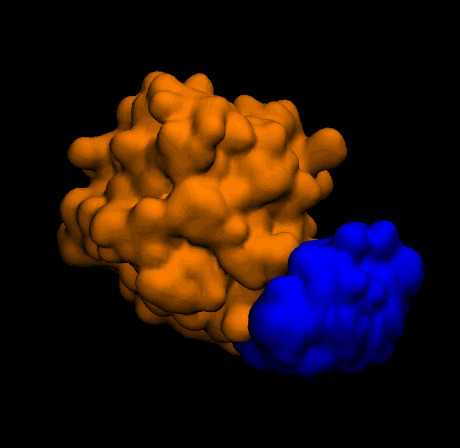
- Q2c. Perform rigid-body protein-ligand docking with Metropolis Monte Carlo using
complex_energy, save accepted poses, and report the final complex energy.
Outputs
Closing thoughts
It has been a while since I looked back on this project and reviewing the papers on protein docking. I was aware of the use of AlphaFold based methods for protein docking but it was nice to see that the Kozakov/Vajda labs use it alongside their physics methods. It seems that the work that has been done to carefully study the interactions of atoms still will be useful when it comes to predicting complexes espcially in the case where there is a lack of information in the dataset.
Enjoy Reading This Article?
Here are some more articles you might like to read next: