A Multistate Kinetic Model of the Sodium-Potassium ATPase
I published a paper with Huan Rui and Benoît Roux in The Journal of Physical Chemistry B titled Multistate Kinetic Model of the Sodium-Potassium ATPase in September 2025. I wanted to write a more informal companion note here because the paper is fairly dense, and the main framework is easier to appreciate in plain language. This paper lays out the theoretical foundation for my dissertation research on modeling P-type ATPase pumps.
How do we turn the familiar Post-Albers cycle into a thermodynamically consistent multistate model that can connect structures, electrophysiology, and atomistic simulation?
At the textbook level, the story of the pump is clear: for each ATP hydrolyzed, the enzyme exports 3 Na+ ions, imports 2 K+ ions, and moves one net positive charge outward. That overall picture is correct, but it hides a great deal of mechanistic detail. Modern structural work has made it possible to identify many intermediates along the transport cycle. Once that structural information is on the table, the obvious question becomes what kind of kinetic model is detailed enough to make use of it without violating thermodynamics. This also complements many kinetic experiments carried out using fluorescence methods and patch-clamp electrophysiology.
Why add so many states?
One motivation for this paper was that the pump is often discussed at two very different scales. At one level, there are measurements of flux and charge movement. At another, there are atomic motions, the diffusion of potassium and sodium into the binding site, the hydrolysis of ATP, and many other events along the cycle.
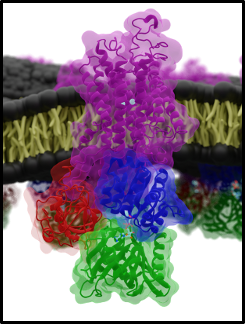
What we tried to build is a bridge between those two perspectives. In the paper, the Na,K-ATPase is represented as a continuous-time Markov model with 20 microstates arranged along the transport cycle. The goal was not to claim that every rate constant is already known precisely. The goal was to set up a framework that respects the right physical constraints and is detailed enough to serve as a landing place for future structural and computational work.
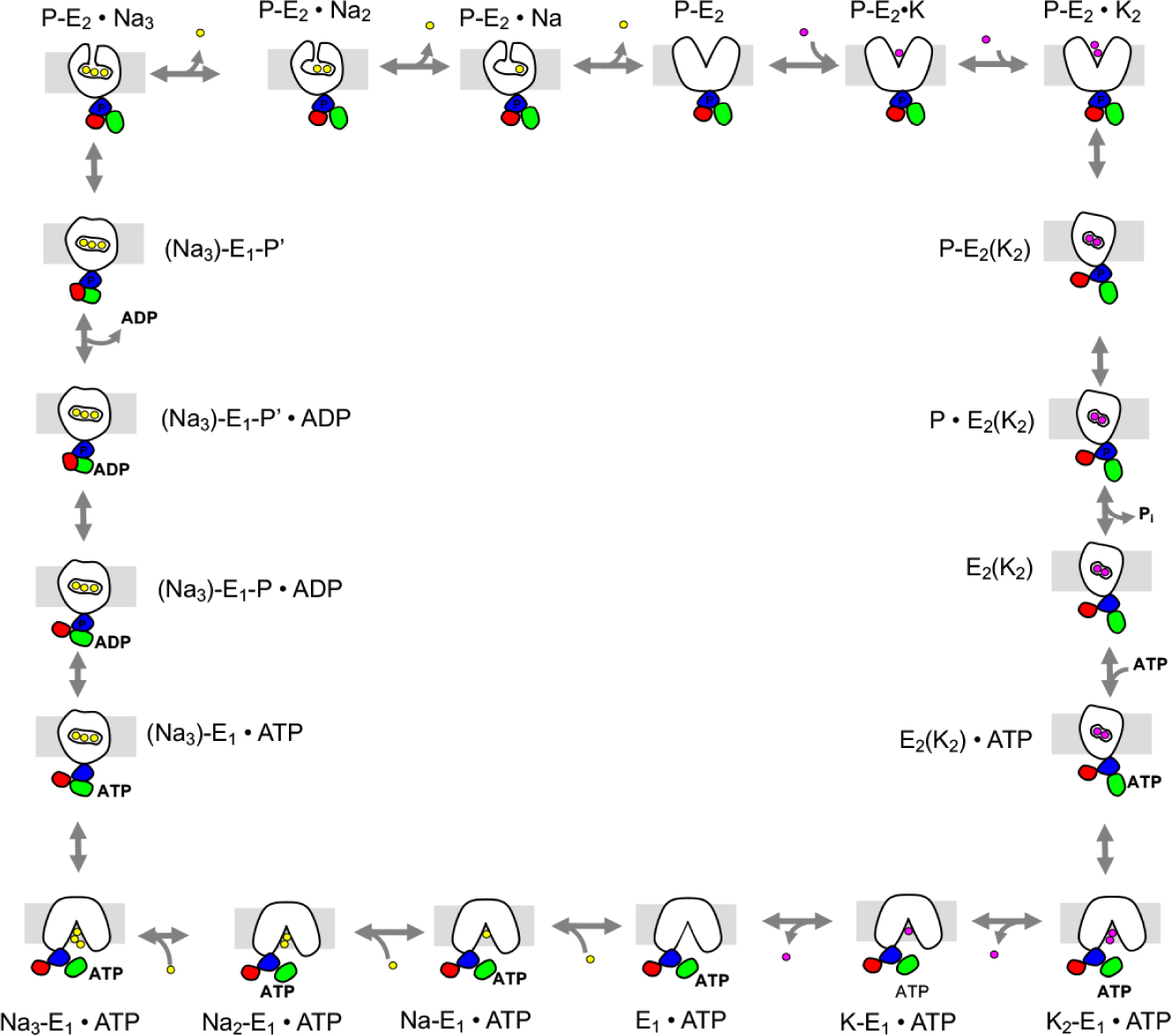
Each of these states corresponds to a specific computational system that can be studied. In fact, we have simulated each one, though that is a topic for another day. The framework allows us to directly connect events that can be investigated computationally with different flavors of molecular dynamics (MD).
What the model actually keeps track of
The model is built so that the microscopic transitions obey the constraints they ought to obey at equilibrium. Specifically, we know that the system is powered by ATP hydrolysis, so we use that as a natural thermodynamic constraint. We also know that the net transfer is +1e, which gives us another natural constraint to impose. Substrate binding and release are incorporated through pseudo-unimolecular rates, but the underlying concentration dependences are still explicit. In other words, ATP, ADP, Pi, Na+, and K+ are not treated as vague background conditions. They are part of the bookkeeping.
At the cycle level, the key quantity is the total free energy change per turnover,
\[\Delta G_{\mathrm{tot}} = -\Delta \mu + W\]where $\Delta \mu$ is the free energy made available by ATP hydrolysis under the cellular conditions of interest and $W$ is the work required to move the ions against their gradients and across the membrane potential. In the paper, using representative physiological conditions for a resting cell, this gives roughly $\Delta \mu = 12$ kcal/mol, $W = 9.9$ kcal/mol, and therefore $\Delta G_{\mathrm{tot}} = -2.1$ kcal/mol. That negative value is what keeps the forward cycle moving.
I think this is an important point. A lot of discussion about ion pumps focuses on stoichiometry, but stoichiometry alone does not tell you how the free energy is partitioned among microscopic steps. That partitioning is exactly what matters if you care about rate limitation, voltage dependence, or functional optimization. A detailed view of these processes can help identify which states are rate limiting and how the landscape changes under different conditions.
Why voltage dependence matters so much
For me, the most interesting part of the paper is the treatment of membrane potential. It is tempting to think of voltage as something that acts only on the overall net transported charge after one full cycle. But that is too coarse. Different microscopic steps can couple to the membrane potential in different ways, and those step-specific couplings can influence the turnover rate strongly.
To make that concrete, we describe the voltage dependence in terms of incremental displacement charges. The idea is that each microscopic transition can carry its own electrogenic signature. This includes not only the transported ions themselves but, in principle, the displacement of charge throughout the whole system: protein, solvent, lipids, and ions. From that point of view, the relevant question is not just “what is the net charge moved per cycle?” but also “how is charge motion distributed across the sequence of intermediate states and barriers?”
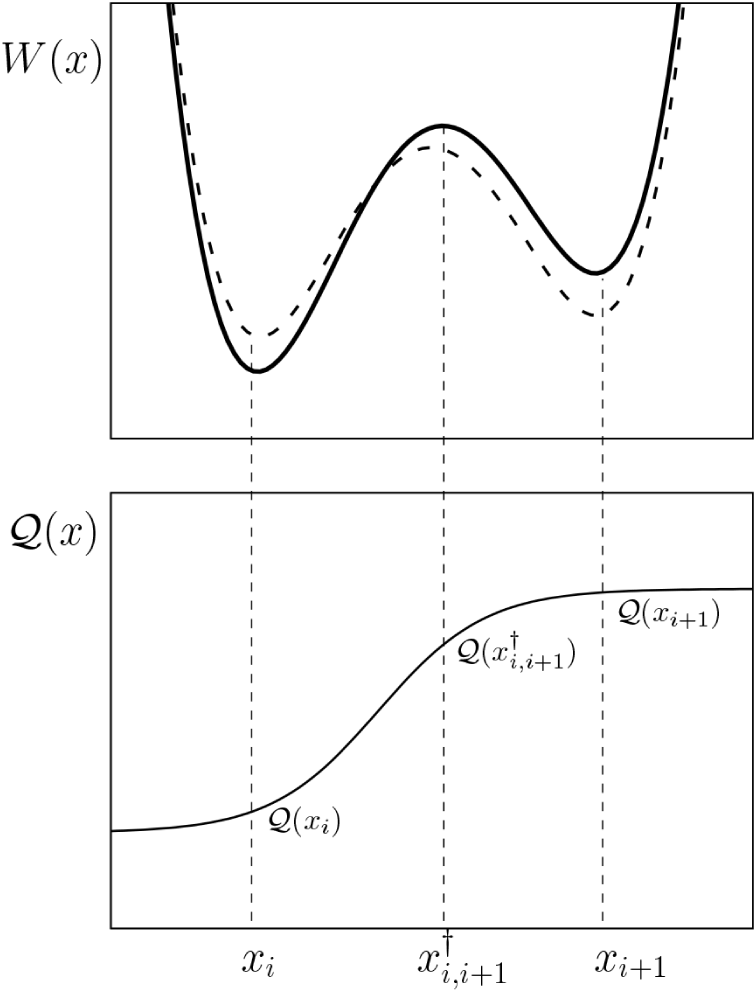
That distinction matters because two transport cycles with the same net stoichiometry do not have to respond to voltage in the same way. The turnover depends on how the free energy barriers and charge increments are allocated over the microscopic steps, not only on the final balance sheet.
In the follow-up to this work, we will present how the incremental charge is distributed across the whole cycle, whereas experimental studies have mainly been able to investigate the voltage dependence of the extracellular half of the cycle. In patch-clamp experiments, people are generally only able to measure the difference in incremental charge between state $i$ and state $i+1$, but computational techniques make it possible to measure $Q(x)$, a continuous function describing the incremental charge. The figure sketches what such an incremental charge function may look like and how it relates to the free energy of the system when a voltage is applied. The point is that it shows how much the free energy changes for a given voltage.
What I think the paper contributes
I do not see this paper as a final fitted model of every elementary step. I see it as a framework paper. Its contribution is that it organizes the problem in a way that future work can refine instead of rebuilding from scratch each time. Concretely, I think it does four useful things:
- It shows how to formulate a 20-state Post-Albers-based kinetic cycle that is compatible with thermodynamic constraints.
- It makes the coupling to membrane potential explicit through state and transition displacement charges rather than treating voltage as an afterthought.
- It creates a natural place where atomistic MD calculations can contribute information that experiments do not yet resolve directly.
- It reframes the pump as a molecular machine whose turnover depends on the full free energy landscape, not just the overall stoichiometry.
We use this as a springboard for the follow-up paper, in which I have simulated all of these states and aim to build a more textbook-like understanding of the pumping cycle at the atomic level.
What the model misses
The current model follows a single main transport cycle. Real pumps are much more complicated. Parallel branches, slippage, and proton leak are all relevant complications, and we discuss that in the paper. In fact, one reason I like this problem is that the Na,K-ATPase is not just a clean schematic object. It is a dynamic membrane protein with multiple ways for the mechanism to become more complicated than the canonical cycle.
That also means the hardest missing information is microscopic. We still need better constraints on the displacement charges associated with individual steps and better estimates of the rate-limiting regions of the cycle. This is exactly where atomistic simulation should become useful, not as a replacement for experiment, but as a complement to it.
Why I think this direction is exciting
What I like most about kinetic models of this kind is that they can become a bridge between scales. Structures tell us what states plausibly exist. Electrophysiology constrains what the pump actually does. Molecular simulation can estimate quantities that are difficult to extract directly from experiment. A multistate kinetic framework is where those pieces can meet. It is also where a theoretical chemist can start to have a direct conversation with an electrophysiologist.
More broadly, I think that this is one of the most interesting features of computational chemistry and biophysics right now. We are no longer forced to choose between a hand-wavy cartoon and a giant atomistic data set. There is room for a middle layer of theory that is detailed enough to be useful and simple enough to reason about. This paper is our attempt to contribute to that middle layer for the Na,K-ATPase. I can imagine that this is the direction future computational biophysics will take. Increases in computational power and the ability to model more complex phenomena make the work I started five years ago more feasible and potentially more useful for understanding the mechanisms of biological processes.
Enjoy Reading This Article?
Here are some more articles you might like to read next: