Bag of Words
This post reviews a homework assignment in which I used a Bag of Words model to analyze 28 scientific papers. By converting each document into a vector representation, I tested whether a simple NLP pipeline could recover meaningful scientific subtopics from the corpus.
Bag of Words is one of the earliest and most influential vector space approaches in natural language processing. While it lacks the sophistication of modern language models, it provides an interpretable foundation for understanding how text can be represented numerically and compared across documents. I present this analysis as an introduction to computational text representation and the broader ideas it motivates.
Overview
1) Preprocessing and vectorization
- lowercase conversion
- punctuation removal
- whitespace token splitting
- term-document matrix construction
From the raw count matrix (X), I computed TF-IDF:
\[T_{d,t} = \frac{X_{d,t}}{\sum_{t'} X_{d,t'}} \cdot \log\left( \frac{N}{ \sum_{d'} \mathbf{1}[X_{d',t} > 0] } \right)\]Then each document vector was L2-normalized.
2) Dimensionality reduction and similarity
- PCA on normalized TF-IDF vectors
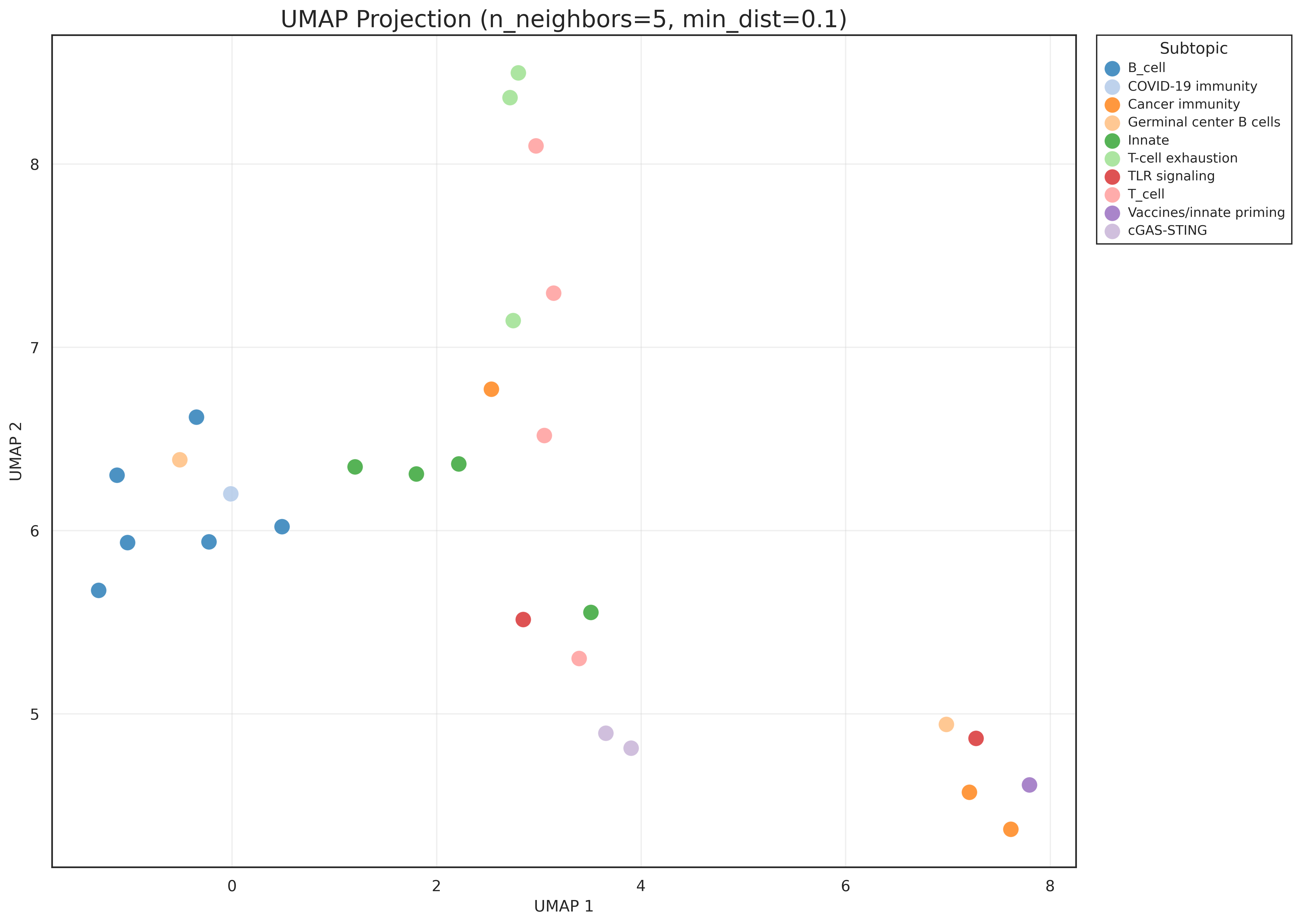
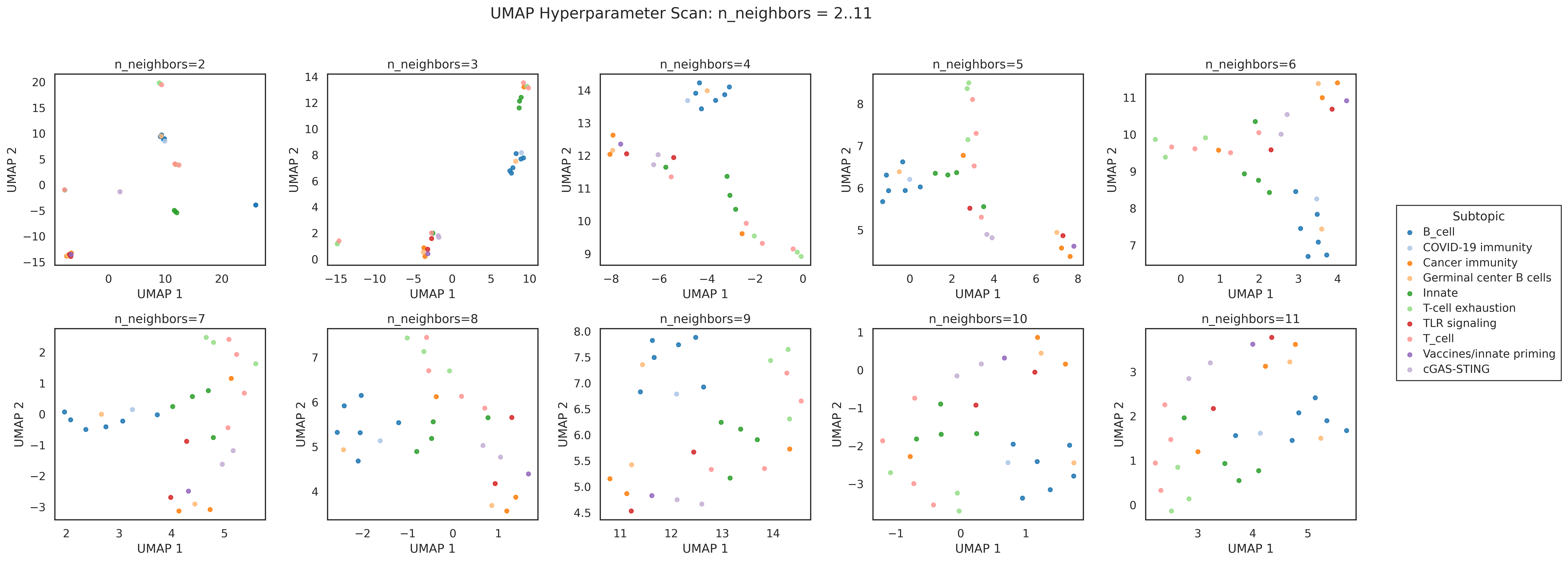
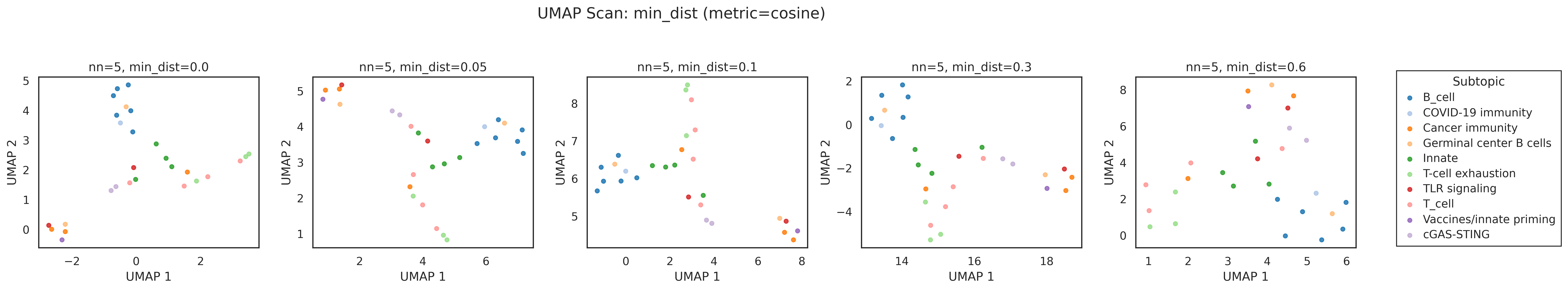
- UMAP with cosine distance (
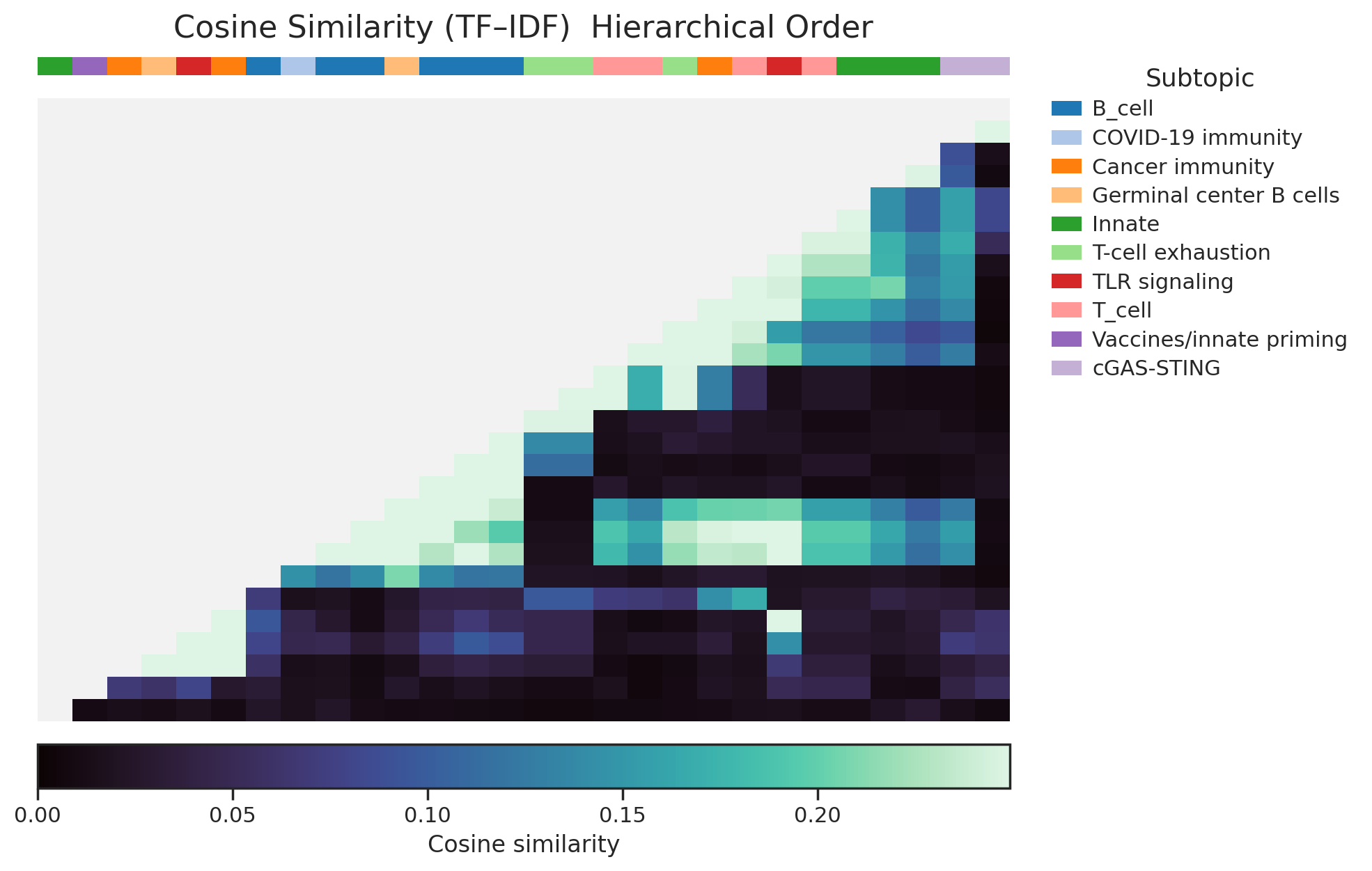
n_neighbors=5,min_dist=0.1) - cosine similarity matrix + hierarchical ordering using distance (1 - \cos(i,j))
Main findings
- papers with similar subtopics tended to group together
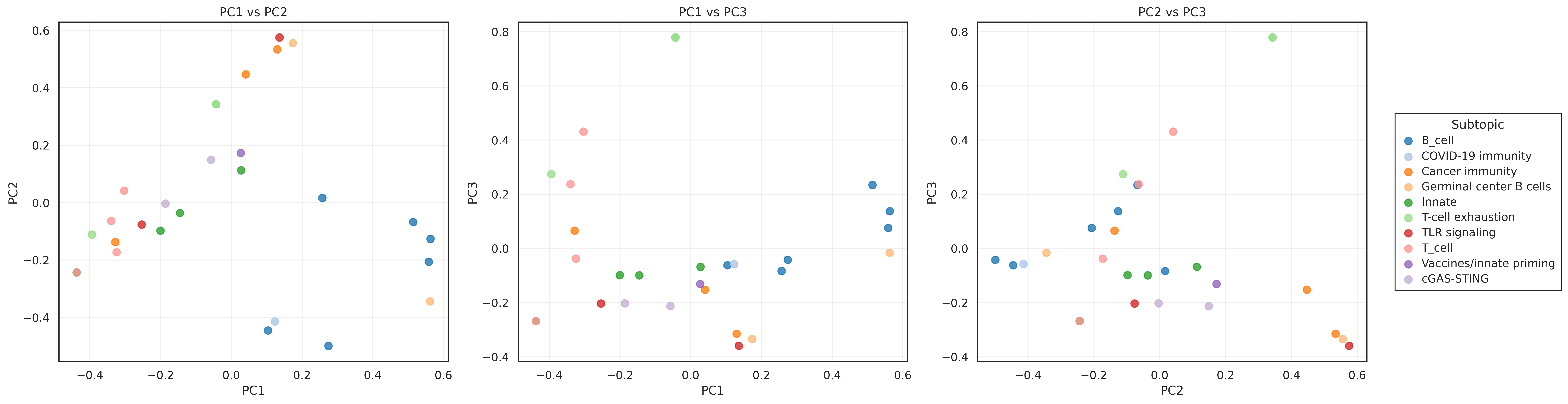
- the first three PCA components explained
0.106,0.100, and0.095of variance - PCA loadings reflected biologically meaningful terms (for example
tfh,gc,germinal,follicular) - UMAP revealed groupings not visible in the PCA projection
- the cosine clustermap was consistent with those grouping patterns
Additional comments
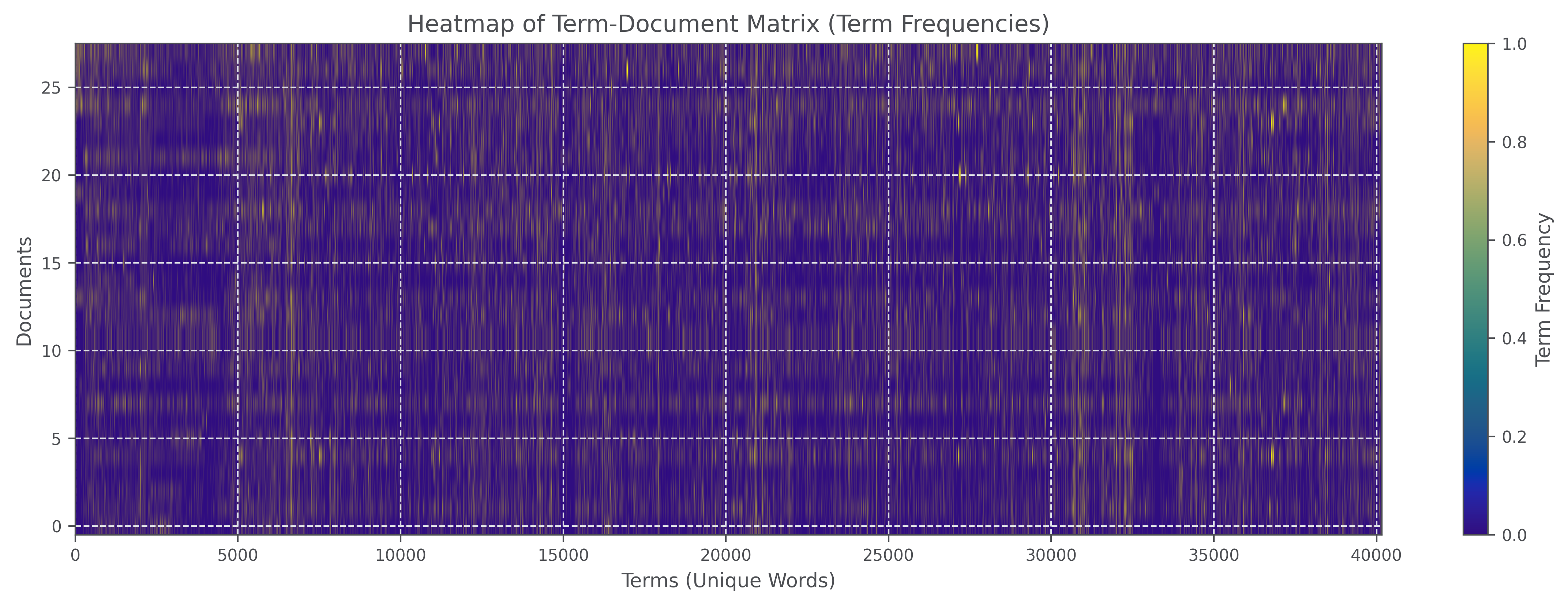
The utility of TF-IDF weighting and L2 normalization was a key part of this assignment. When I built the raw term-document matrix, the most frequent token was, unsurprisingly, and.
This illustrates why raw counts alone can produce weak embeddings: high-frequency background tokens dominate the signal across nearly all documents. After TF-IDF weighting, high-weight terms shifted toward more domain-specific vocabulary such as cgas.
| Rank | Term | Count |
|---|---|---|
| 1 | and | 14573 |
| 2 | of | 13337 |
| 3 | the | 12363 |
| 4 | in | 10033 |
| 5 | a | 6960 |
| 6 | to | 6528 |
| 7 | cells | 6462 |
| 8 | t | 6120 |
| 9 | al | 5411 |
| 10 | et | 5343 |
The token t (rank 8) is likely an artifact of PDF extraction splitting hyphenated terms such as “T-cell” into t and cell.
Nearest-neighbor check
| Query document | Doc 0 |
|---|---|
| Nearest neighbor | Doc 6 |
| Cosine similarity | 0.6177 |
| Doc | Title | Subtopic |
|---|---|---|
| 0 | Molecular and cellular insights into T cell exhaustion. | T-cell exhaustion |
| 6 | Defining "T cell exhaustion". | T-cell exhaustion |
| Term | Shared score |
|---|---|
| exhausted | 0.2281 |
| exhaustion | 0.1415 |
| pd1 | 0.0550 |
| pubmed | 0.0462 |
| manuscript | 0.0460 |
| author | 0.0364 |
| wherry | 0.0048 |
| tumour | 0.0042 |
| pmc | 0.0035 |
| progenitor | 0.0029 |
| tumours | 0.0027 |
| cd8 | 0.0020 |
Several top shared terms (pubmed, manuscript, author, pmc) are metadata artifacts from PDF extraction. The cosine similarity of 0.6177 is therefore partially inflated by source noise rather than topic signal alone.
Supplementary: top PCA loading tables (PC1 / PC2 / PC3)
| Term | Loading |
|---|---|
| gc | 0.4543 |
| tfh | 0.2394 |
| germinal | 0.1579 |
| org | 0.1191 |
| gcs | 0.1136 |
| lz | 0.1119 |
| follicular | 0.1003 |
| annualreviews | 0.0984 |
| dz | 0.0959 |
| https | 0.0869 |
| self-reactive | 0.0822 |
| pubmed | -0.2997 |
| manuscript | -0.2147 |
| autophagy | -0.2022 |
| author | -0.1893 |
| exhausted | -0.1622 |
| exhaustion | -0.1589 |
| deretic | -0.1132 |
| tex | -0.0924 |
| pd1 | -0.0754 |
| Term | Loading |
|---|---|
| https | 0.3579 |
| org | 0.3322 |
| tex | 0.1823 |
| ammasome | 0.1255 |
| 1038 | 0.0911 |
| ammatory | 0.0862 |
| guest | 0.0773 |
| gc | -0.3272 |
| pubmed | -0.3113 |
| manuscript | -0.2155 |
| author | -0.1884 |
| germinal | -0.1300 |
| autophagy | -0.1139 |
| gcs | -0.1081 |
| dz | -0.1043 |
| lz | -0.1042 |
| tfh | -0.0774 |
| shm | -0.0714 |
| deretic | -0.0667 |
| bcr | -0.0653 |
Note: ammasome and ammatory appear to be truncated tokens from PDF extraction, likely fragments of inflammasome and inflammatory.
| Term | Loading |
|---|---|
| tex | 0.5089 |
| exhaustion | 0.3057 |
| guest | 0.2314 |
| exhausted | 0.1664 |
| annualreviews | 0.1507 |
| iy37ch19-wherry | 0.1199 |
| pd-1 | 0.1158 |
| cls | 0.1157 |
| arjats | 0.1157 |
| etal | 0.1138 |
| tmem | 0.1073 |
| teff | 0.1057 |
| downloaded | 0.0997 |
| 2026 | 0.0997 |
| www | 0.0970 |
| mon | 0.0907 |
| https | -0.2133 |
| autophagy | -0.1412 |
| org | -0.1320 |
| cgas | -0.1191 |
Limitations
- preprocessing was intentionally simple, so metadata/noise leaked into features
- tokens such as
annualreviews,https,pubmed, andauthorappeared with large weights - BoW/TF-IDF ignores word order and deeper semantics
Takeaway
Bag-of-Words + TF-IDF provided an interpretable baseline that partially recovered subtopic structure in this 28-paper corpus, though the results were strongly affected by simple preprocessing choices and residual PDF/source metadata.
Enjoy Reading This Article?
Here are some more articles you might like to read next: